Cell Communication and Migration Laboratory (also CellComM, abbreviated CCM, or Sáez Lab).
We are a multidisciplinary team, which goal is to understand the molecular mechanisms underlying cell migration and communication across different scales.
Cell migration is key for different phenomenon such as development, tissue reparation, the immune response, and cancer metastasis. Importantly, cells migrate in a confined microenvironment, which is composed of extracellular matrix as well as other cells. The coordination of the migratory response is achieved by both cell autonomous signaling, and intercellular communication. During collective cell migration, cell-to-cell communication plays a major role. On the other hand, if there is coordination of single migrating cells is much less understood.
Projects
Project 1
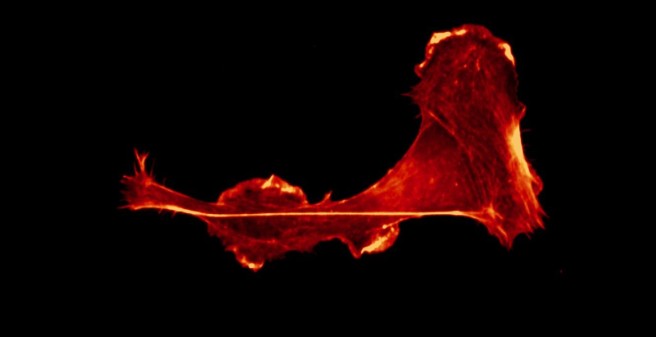
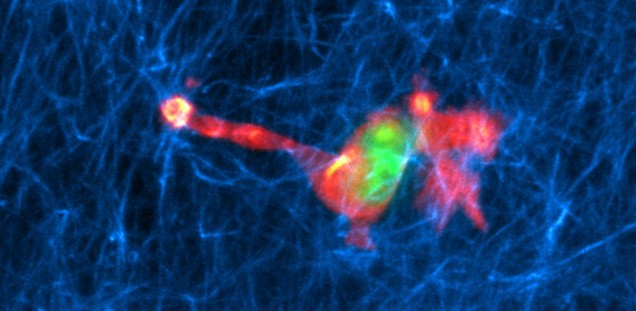
Immune cell migration. Leukocytes, such as dendritic cells and T cells, constantly patrol our body in search for danger signals. These danger signals can be associated to pathogens (pathogen-associated molecular patters, PAMPs) or to cell damage (damage-associated molecular patterns, DAMPs). Interestingly, both PAMPs (i.e. bacterial LPS) and DAMPs (i.e. ATP) trigger fast migration of immune cells, although the signaling is different. This is explained by signaling convergence at the level of the cytoskeleton, because in both cases fast migrating dendritic cells present a re-localization of the F-actin pool towards the cell rear, where contractility occurs. Moreover, the calcium signaling triggered by LPS is predominantly from intracellular stores, including the lysosomes ( PMID 29079589 ), whereas the one induced by ATP relies on Ca2+ influx and autocrine ATP release via the P2X7-Panx1 axis ( PMID 29162744 ). At the CellComM lab we use microfluidics, 3D culture, artificial extracellular matrices, and live-cell imaging to understand how leukocytes migrate efficiently during chemotaxis. In the context of the SFB1328 (project A20) we aim to unveil the role of ATP and other adenine nucleotides during immune cell migration.
Project 2
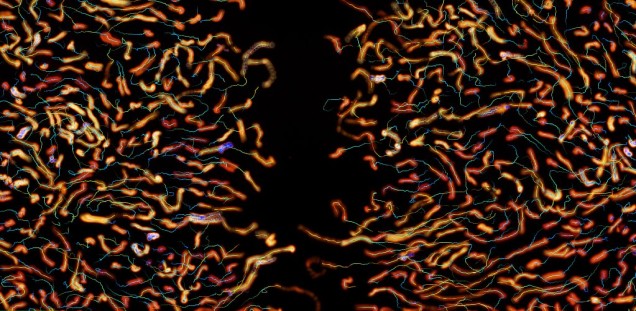
Collective cell migration. Intercellular coordination is key during collective cell migration. In particular, cell communication and migration are tightly regulated during angiogenesis, the formation of new vessels. The physiology of endothelial cells, which form those blood vessels, is directly affected in different diseases, such as obesity (
PMID 30006153
). In collaboration with physicists and mathematicians, at the CellComM lab we combine the visualization of cell behavior using microscopy with theoretical analysis. The goal is to develop new models to predict cellular interactions (
PMID 32195344
), and behaviour based on migratory responses at the cellular level. In addition, we are also interested in the directional cell migration of single cells, which normally migrate collectively (
bioRxiv
). In the context of a Human Frontiers Scientific Program (HFSP) grant we aim to understand how cells decide spontaneously for a new direction.
Project 3
Interorganelle crosstalk. Communication and interaction occur not only at the cellular scale, but also at the subcellular one. The coordination and crosstalk between organelles has emerged as key for a plethora of responses. At the CellComM lab we aim to determine how the interaction between different organelles control cell fate. In collaboration with the AG Heeren, and in the context of the ITN EndoConnect , we aim to understand the role of endolysosomal signaling in adipocyte function and other cellular models. Using state-of-art microscopy and image analysis tools developed in our lab in combination with the toolkit available in the ITN EndoConnect consortium, we are exploring the interactions of the subcellular world.
For a full publication list of the PI: https://scholar.google.com/citations
For a full publication list of the CCM lab: https://scholar.google.com/citations
Collaborations
Yamuna Krishnan, Chicago University, US
Nir Gov, Weizmann Institute Rehovot Israel
Matthieu Piel, Ana-Maria Lennon, Institut Curie, Paris
Maria Teresa Alonso, Instituto De Biología Y Genética Molecular, Valladolid, Spain
Silvana Zanlungo, CIM, PUC, Chile
Paloma Harcha, Adrian Palacios, Juan Carlos Sáez, Centro de Neurociencia Valparaíso, Valpo, Chile Nicolas Demaurex, Dept of Cell Physiology and Metabolism, Université de Genève, Genève
Sophie Acton, Medical Research Council Laboratory for Molecular Cell Biology, U. College London, UK
Nils Gauthier Instituto Firc Di Oncologia Molecolare, Milano, Italy
Christian Grimm, Ludwig-Maximilians-Univ (LMU) Dpt Pharmacology and Toxicology München, Germany
Catalina Lodillinsky, Institute of Oncology Angel H. Roffo, Buenos Aires, Argentina
Judith Klumperman, Ph.D. Department of Cell Biology | Center for Molecular Medicine | University Medical Center Utrecht
Baldur Schroeter, Institut für Thermische Verfahrenstechnik, Technische Universität Hamburg
Maike Frye, Institut für Klinische Chemie und Laboratoriumsmedizin, UKE, Hamburg
Nicola Gagliani, SFB1328, UKE, Hamburg
Jörg Heeren, SFB1328, UKE, Hamburg