 UKE homepage?
UKE homepage?

Multidimensional analysis of protective and damaging adaptive immune responses in emerging viral infections
Summary
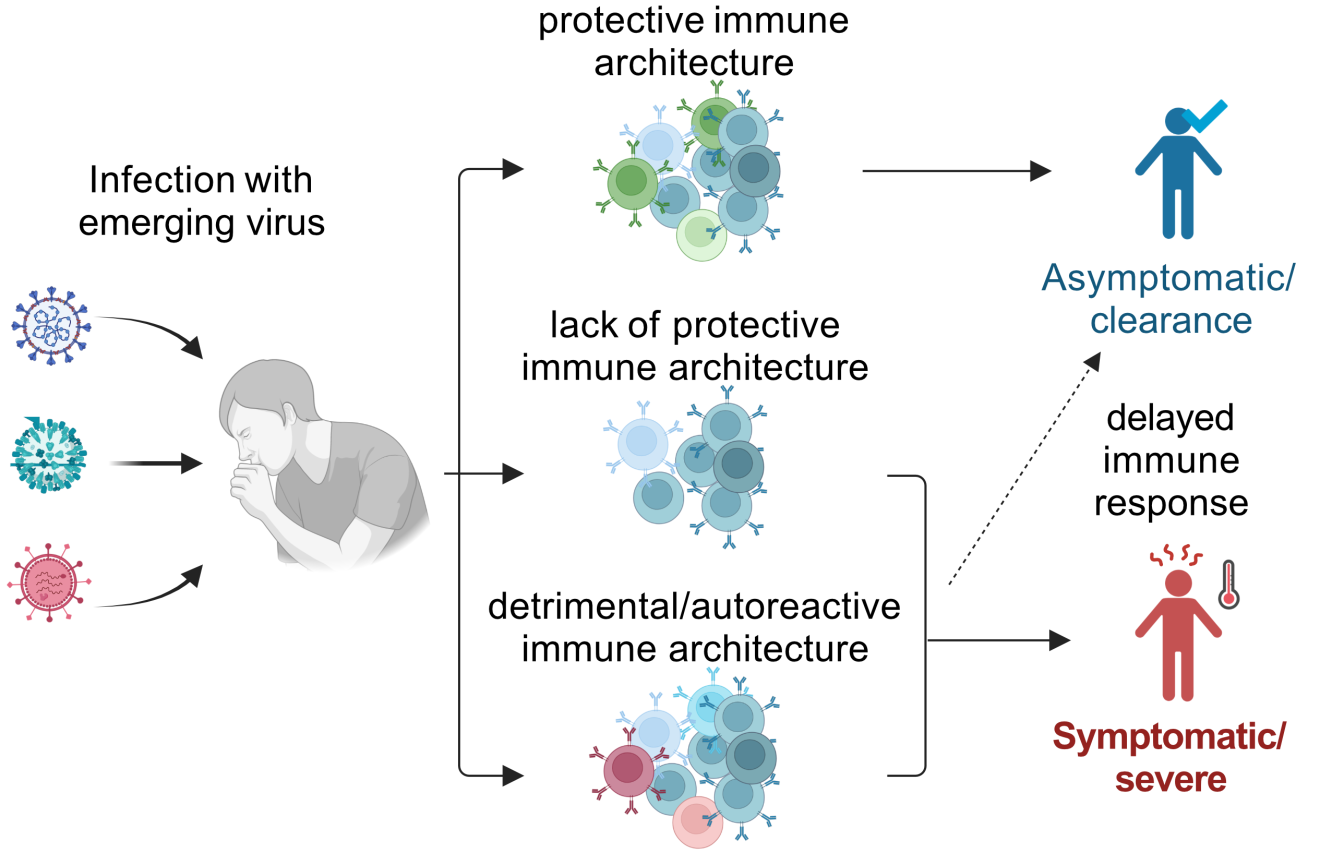
Emerging and re-emerging viruses - like SARS-CoV-2, Mpox, or avian influenza - pose ongoing risks to global health. In response, it's crucial to understand how the immune system reacts to these new threats, especially in vulnerable groups. Initial immune responses to novel pathogens are often slower and less efficient than responses to known viruses, which can allow the infection to spread and trigger harmful inflammation.
This project explores how early adaptive immune responses contribute both to clearing the virus and to potential immunopathology, such as hyperinflammation or autoimmune-like effects. By identifying shared immune response patterns across different viruses and patient groups, we aim to improve risk assessment and guide the development of targeted vaccines and therapies - especially in immunocompromised individuals or those with repeat infections.
Our Work Packages (WP):
WP1: Multidimensional profiling of B and T cell repertoires, interactomes, and phenotypes in COVID-19 and Mpox patients
We investigate B and T cell responses in SARS-CoV-2 and Mpox patients through comprehensive immune profiling to identify patterns that distinguish protective responses from detrimental ones, aiming to generate a detailed immune fingerprint applicable to other viral infections.
WP2: ML-based identification of multidimensional correlates of adaptive immunity during protective, damaging, and persisting SARS-CoV-2 infection
We use machine learning to analyze immune data from WP1, aiming to identify patterns that correlate with either protective immunity or harmful immune responses, enhancing predictive tools for patient outcomes in viral infections.
WP3: Validation of human immunoreceptor network-derived SARS-CoV-2 antibodies
We validate antibodies derived from specific immune receptor networks to assess their reactivity to SARS-CoV-2, helping to understand immune responses linked to either effective viral clearance or severe disease.
With these work packages, we aim to better understand the immune responses to viral infections and use this knowledge to improve diagnostics, predict outcomes, and develop targeted therapies for both current and future pandemics.
Team


Prof. Dr. Mascha Binder
Principal Investigator
E-mail address:


Prof. Dr. Julian Schulze zur Wiesch
Principal Investigator
E-mail address:
Phone: +49 (0) 40 7410 - 52831, +49 (0) 40 7410 - 20977




Research Group Binder
The Research Group Translational Immuno-Oncology is based at the University of Basel.
Research Group Schulze zur Wiesch
The Research Group Role of T cells in infectious diseases is based at the University Medical Center Hamburg-Eppendorf (UKE).
Project related publications
von der Schulenburg P, Behrens GMN, Hoffmann M, Linke A, Nehlmeier I, Kempf AM, Stankov M, Lütgehetmann M, Jahnke-Triankowski J, Addo MM, Fischer L, Lohse AW, Pöhlmann S, Schulze Zur Wiesch J, Sterneck M. Immune Response to SARS-CoV-2 XBB.1.5 and JN.1 Variants Following XBB.1.5 Booster Vaccination in Liver Transplant Recipients. Viruses. 2024 Dec 19;16(12):1942. doi: 10.3390/v16121942.
Jensen BEO#, Knops E#, Cords L#, Lübke N#, Salgado M#, …, Schulze zur Wiesch J#, Wensing AMJ#, Martinez-Picado J#, Kobbe G#. In-depth virological and immunological characterization of HIV-1 cure after CCR5Δ32/Δ32 allogeneic hematopoietic stem cell transplantation. Nat Med. 2023;29(3):583-587. doi: 10.1038/s41591-023-02213-x.
Schultheiß C, Willscher E, Paschold L, …, Gekle M, Mikolajczyk R, Binder M. The IL-1ß, IL-6 and TNF cytokine triad is associated with post-acute sequelae of COVID-19. Cell Rep Med. 2022;3(6):100663. doi: 10.1016/j.xcrm.2022.100663.
Zhao Y#, Kilian C#, Turner JE#, Bosurgi L#, Roedl K#, …, Schulze zur Wiesch J, …, Addo MM, Lohse AW, Binder M, …, Panzer U#, Gagliani N#, Krebs CF#. Clonal expansion and activation of tissue-resident memory-like Th17 cells expressing GM-CSF in the lung of severe COVID-19 patients. Sci Immunol. 2021;6(56):eabf6692. doi: 10.1126/sciimmunol.abf6692.
Heide J, Schulte S, Kohsar M, …, Kwok WW, Sidney J, Schulze Zur Wiesch J. Broadly directed SARS-CoV-2-specific CD4+ T cell response includes frequently detected peptide specificities within the membrane and nucleoprotein in patients with acute and resolved COVID-19. PLoS Pathog. 2021;17(9):e1009842. doi: 10.1371/journal.ppat.1009842.
Simnica D#, Schultheiß C#, Mohme M#, …, Gagliani N, …, Heide J, Schulze-zur-Wiesch J, Binder M. Landscape of T cell repertoires with public COVID-19-associated T cell receptors in pre-pandemic risk cohorts. Clin Transl Immunol. 2021;10(9):e1340. doi: 10.1002/cti2.
Schultheiß C, Paschold L, Willscher E, …, Keysser G, Gagliani N, Binder M. Maturation trajectories and transcriptional landscape of plasmablasts and autoreactive B cells in COVID-19. iScience. 2021;24(11):103325. doi: 10.1016/j.isci.2021.103325.
Paschold L#, Simnica D#, Willscher E, …, Sedding DG, Schultheiß C, Binder M. SARS-CoV-2 specific antibody rearrangements in pre-pandemic immune repertoires of risk cohorts and COVID-19 patients. J Clin Invest. 2021;131(1):e142966. doi: 10.1172/JCI142966.
Wildner NH, Ahmadi P, Schulte S, …, Addo MM, Haag F, Schulze zur Wiesch J. B cell analysis in SARS-CoV-2 versus malaria: Increased frequencies of plasmablasts and atypical memory B cells in COVID-19. J Leukoc Biol. 2021;109(1):77-90. doi: 10.1002/JLB.5COVA0620-370RR.
Schultheiß C#, Paschold L#, Simnica D#, …, Ciesek S, Addo M, Binder M. Next Generation Sequencing of T and B cell receptor repertoires from COVID-19 patients showed signatures associated with severity of disease. Immunity. 2020;53(2):442-455. doi: 10.1016/j.immuni.2020.06.024.
Cheng MH, Zhang S, Porritt RA, …, Binder M, Arditi M#, Bahar I#. Superantigenic character of an insert unique to SARS-CoV-2 spike supported by skewed TCR repertoire in patients with hyperinflammation. Proc Natl Acad Sci USA. 2020;117(41):25254-25262. doi: 10.1073/pnas.2010722117.
#equally contributing authors