 UKE homepage?
UKE homepage?

Determinants of heterospecific phase separation in negative-strand RNA viruses
Summary
Negative-strand RNA viruses (NSVs) replicate inside cells in specialized structures called inclusion bodies (IBs). These are membrane-less compartments formed by viral and host molecules, often through a process known as liquid-liquid phase separation (LLPS). Although many proteins have the potential to form such structures, it is still unclear why only certain ones are recruited into viral IBs — and what role the viral genome plays in this process. We aim to uncover the underlying rules that guide the formation of these viral compartments. Understanding these principles could help us design new inhibitors that may be used to combat both known viruses and future threats like "Disease X."
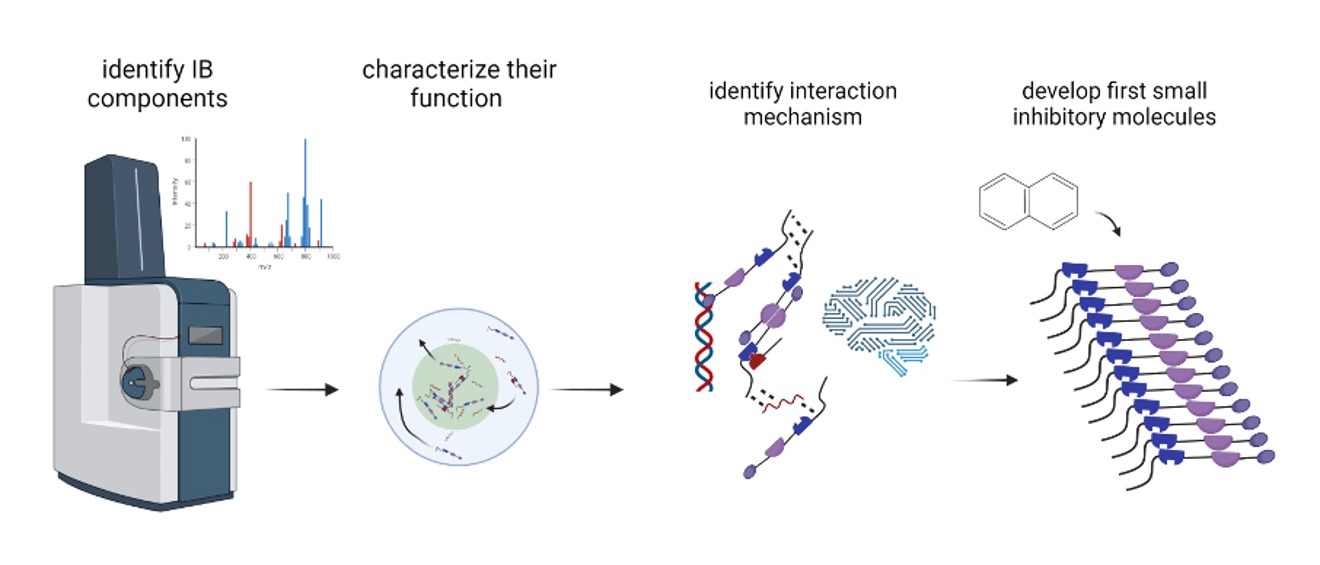
Our Work Packages (WP):
WP1: Enabling Click-labeling of Viral RNA in Living Cells Using the TriPPPro-technology - we develop a new labeling method to visualize viral RNA in living cells and track where and when it interacts with other molecules during infection.
WP2: Identification of Proteins Residing in Genome-Associated Inclusion Bodies - we identify which cellular and viral proteins are present in viral inclusion bodies and study how they are connected to viral RNA and replication.
WP3: Deciphering the Mechanism of Recruitment into Phase-Separated Inclusion Bodies - we investigate how specific proteins are selectively recruited into inclusion bodies and which parts of their structure make this possible.
WP4: Designing IB-Specific Inhibitors by Targeting Condensate-Specific Interactions - we use the insights gained from WP1–3 to design small molecules that disrupt or stabilize viral inclusion bodies and block viral replication.
These work packages help us understand how viral inclusion bodies (IBs) form and function - and how we might target them to stop the spread of dangerous viruses.
Team


Prof. Dr. Chris Meier
Principal Investigator
E-mail address:
Phone: +49 40 42838-4324


Prof. Dr. Jens Bosse
Principal Investigator
E-mail address:
Phone: +49 040 8998 87645





Research Group Meier
The Research Group Synthetic Organic Chemistry, Medicinal Chemistry and Chemical Biology is based at the University of Hamburg.
Research Group Bosse
The Research Group Quantitative Virology is based at the Centre for Structural Systems Biology (CSSB).
Project related publications
Sterrenberg VT#, Stalling D#, Knaack JIH, Soh TK, Bosse JB#, Meier C#. A TriPPPro-Nucleotide Reporter with Optimized Cell-Permeable Dyes for Metabolic Labeling of Cellular and Viral DNA in Living Cells. Angew Chem Int Ed Engl. 2023;62(38):e202308271. doi: 10.1002/anie.202308271.
Caragliano E, Bonazza S, Frascaroli G, …, Grünewald K, Bosse JB#, Brune W#. Human cytomegalovirus forms phase-separated compartments at viral genomes to facilitate viral replication. Cell Rep. 2022;38(10):110469. doi: 10.1016/j.celrep.2022.110469.
Zhao C, Jia X, Schols D, Balzarini J, Meier C. gamma-Non-Symmetrically Dimasked TriPPPro Prodrugs as Potential Antiviral Agents against HIV. ChemMedChem. 2021;16(3):499–512. doi: 10.1002/cmdc.202000712.
Jung J, Ching W, Baumdick M, …, Bosse JB, …, Belderbos M, Dobner T, Bunders M. KIR3DS1 directs NK cell-mediated protection against human adenovirus infections. Sci Immunol. 2021;6(63):eabe2942. doi: 10.1126/sciimmunol.abe2942.
Jia X, Weber S, Schols D, Balzarini J, Meier C. Membrane Permeable, Bioreversibly Modified Prodrugs of Nucleoside Diphosphate-γ-Phosphonates. J Med Chem. 2020;63(20):11990–12007. doi: 10.1021/acs.jmedchem.0c01294.
Zhao C, Weber S, Schols D, …, Balzarini J, Meier C. Prodrugs of γ-Alkyl-Modified Nucleoside Triphosphates: Improved Inhibition of HIV Reverse Transcriptase. Angew Chem Int Ed Engl. 2020;59(49):22063–22071. doi: 10.1002/anie.202003073.
Gollnest T, Dinis de Oliveira T, Rath AK, …, Schols D, Balzarini J, Meier C. Membrane-permeable Triphosphate Prodrugs of Nucleoside Analogues. Angew Chem Int Ed Engl. 2016;55(17):5255–5258. doi: 10.1002/anie.201511808.
Gollnest T, Dinis de Oliveira T, Schols D, Balzarini J, Meier C. Lipophilic prodrugs of nucleoside triphosphates as biochemical probes and potential antivirals. Nat Commun. 2015;6:8716. doi: 10.1038/ncomms9716.
Bosse JB, Hogue IB, Feric M, …, Sodeik B, Brangwynne CP, Enquist LW. Remodeling nuclear architecture allows efficient transport of herpesvirus capsids by diffusion. Proc Natl Acad Sci U S A. 2015;112(42):E5725–E5733. doi: 10.1073/pnas.1513876112.
Hagen C#, Dent KC#, Zeev-Ben-Mordehai T#, Grange M#, Bosse JB#, …, Plitzko JM, Mettenleiter TC, Grünewald K. Structural Basis of Vesicle Formation at the Inner Nuclear Membrane. Cell. 2015;163(7):1692–1701. doi: 10.1016/j.cell.2015.11.029.
#equally contributing authors