 UKE homepage?
UKE homepage?

Structural cell biology of Bunyaviruses
Summary
Bunyaviruses are globally distributed and can infect a wide variety of organisms. Some members of this virus family, such as Lassa virus (LASV) and Rift Valley fever virus (RVFV), are responsible for recurring outbreaks and are considered a serious threat to public health, as acknowledged by their inclusion on the WHO's R&D Blueprint list. Despite this, the biology of bunyaviruses remains poorly understood. These viruses are composed of small, membrane-bound particles and rely on a minimal set of viral proteins - often multifunctional - to complete their life cycle. One key example is the L protein, a large and complex enzyme responsible for copying and transcribing the viral genome.
Although some structural details of the L protein have been determined, much remains unknown about how genome replication and transcription take place inside infected cells, and which host and viral factors are involved. This project aims to close these knowledge gaps by examining how the replication and transcription processes are organized within cells, from virus entry to assembly. Using advanced imaging techniques such as electron cryo-tomography (cryoET) and single-particle cryo-electron microscopy (cryoEM), the researchers hope to uncover critical molecular interactions that could serve as new targets for antiviral drug development.
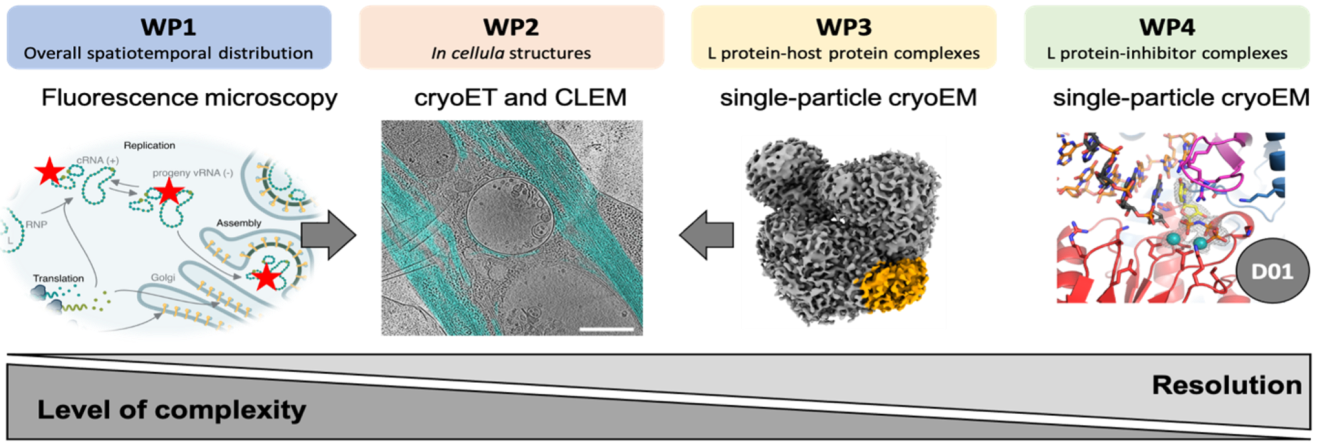
Our Work Packages (WP):
This project is composed of four interconnected work packages that aim to reveal the bunyavirus life cycle at different levels of detail - from spatial localization within cells to high-resolution structural analyses.
WP1: Visualizing viral replication in space and time - we investigate where and when the L protein of LASV and RVFV performs viral transcription and replication in host cells using advanced fluorescence microscopy.
WP2: Structural insights into viral processes in cells - we use cryo-electron tomography (cryoET) and correlative microscopy to visualize how bunyaviruses reshape cellular structures during entry, replication, and egress.
WP3: High-resolution structures of viral polymerase-host interactions - we determine 3D structures of the L protein in complex with host factors and RNA using single-particle cryoEM to understand host-virus interplay at atomic resolution.
WP4: Structural basis of L protein inhibition - we assess the impact of nucleotide inhibitors on the L protein using biochemical assays and high-resolution cryoEM to guide the optimization of antiviral compounds.
These work packages help us to systematically dissect viral replication, host interactions, and inhibitor effects, providing the structural and functional basis for future therapeutic strategies.
Team


Prof. Dr. Kay Grünewald
Principal Investigator
E-mail address:
Phone: +49 40 8998 87700


Prof. Dr. Maria Rosenthal
Principal Investigator
E-mail adress:
Phone: +49 40 285380-930




Department Grünewald
The Department Structural Cell Biology of Viruses is based at the Centre for Structural Systems Biology (CSSB).
Research Group Rosenthal
The Research Group Structural Virology is based at the Bernhard Nocht Institute for Tropical Medicine (BNITM).
Project related publications
Williams HM, Thorkelsson SR, Vogel D, Milewski M, Busch C, Cusack S, Grünewald K, Quemin ERJ, Rosenthal M. Structural insights into viral genome replication by the severe fever with thrombocytopenia syndrome virus L protein. Nucleic Acids Res. 2023;51(3):1424–1442. doi: 10.1093/nar/gkac1249.
Kouba T#, Vogel D#, Thorkelsson SR#, Quemin ERJ, Williams HM, Milewski M, Busch C, Günther S, Grünewald K, Rosenthal M, Cusack S. Conformational changes in Lassa virus L protein associated with promoter binding and RNA synthesis activity. Nat Commun. 2021;12(1):7018. doi: 10.1038/s41467-021-27305-5.
Vogel D, Rosenthal M, Gogrefe N, Reindl S, Günther S. Biochemical characterization of the Lassa virus L protein. J Biol Chem. 2019;294(20):8088. doi: 10.1074/jbc.RA118.006973.
Jérôme H, Rudolf M, Lelke M, Pahlmann M, Busch C, Bockholt S, Wurr S, Günther S, Rosenthal M, Kerber R. Rift Valley fever virus minigenome system for investigating the role of L protein residues in viral transcription and replication. J Gen Virol. 2019;100(7):1093–1098. doi: 10.1099/jgv.0.001281.
Moser F#, Pražák V#, Mordhorst V, Andrade DM, Baker LA, Hagen C, Grünewald K, Kaufmann R. Cryo-SOFI enabling low-dose super-resolution correlative light and electron cryo-microscopy. PNAS. 2019;116(11):4804–4809. doi: 10.1073/pnas.1810690116.
Hagen C#, Dent KC#, Zeev-Ben-Mordehai T#, Grange M#, Bosse JB#, Whittle C, Klupp BG, Siebert CA, Vasishtan D, Bäuerlein FJB, Cheleski J, Werner S, Guttman P, Rehbein S, Henzler K, Demmerle J, Adler B, Koszinowski U, Schermelleh L, Schneider G, Enquist LW, Plitzko JM, Mettenleiter TC, Grünewald K. Structural Basis of Vesicle Formation at the Inner Nuclear Membrane. Cell. 2015;163(7):1692–1701. doi: 10.1016/j.cell.2015.11.029.
Schellenberger P, Kaufmann R, Siebert CA, Hagen C, Wodrich H, Grünewald K. High-precision correlative fluorescence and electron cryo microscopy using two independent alignment markers. Ultramicroscopy. 2014;143(100):41–51. doi: 10.1016/j.ultramic.2013.10.011.
Huiskonen JT, Hepojoki J, Laurinmäki P, Vaheri A, Lankinen H, Butcher SJ, Grünewald K. Electron cryotomography of Tula hantavirus suggests a unique assembly paradigm for enveloped viruses. J Virol. 2010;84(10):4889–4897. doi: 10.1128/JVI.00057-10.
Huiskonen JT, Overby AK, Weber F, Grünewald K. Electron cryo-microscopy and single-particle averaging of Rift Valley fever virus: evidence for GN-GC glycoprotein heterodimers. J Virol. 2009;83(8):3762–3769. doi: 10.1128/JVI.02483-08.
#equally contributing authors